Overview¶
Extracellular recordings are an essential source of data in experimental and clinical neuroscience. Of particular interest in these recordings is the activity of single neurons which must be inferred using a blind source separation procedure called spike sorting.
Given the importance of spike sorting, much attention has been directed towards the development of tools and algorithms that can increase its performance and automation. These developments, however, introduce new challenges in software and file format incompatibility which reduce interoperability, hinder benchmarking, and preclude reproducible analysis.
To address these limitations, we developed SpikeInterface, a Python framework designed to unify preexisting spike sorting technologies into a single code base and to standardize extracellular data file handling. With a few lines of code, users can run, compare, and benchmark most modern spike sorting algorithms; pre-process, post-process, and visualize extracellular datasets; validate, curate, and export sorted results; and more, regardless of the underlying data format.
In the following documentation, we provide an overview of SpikeInterface.
Organization¶
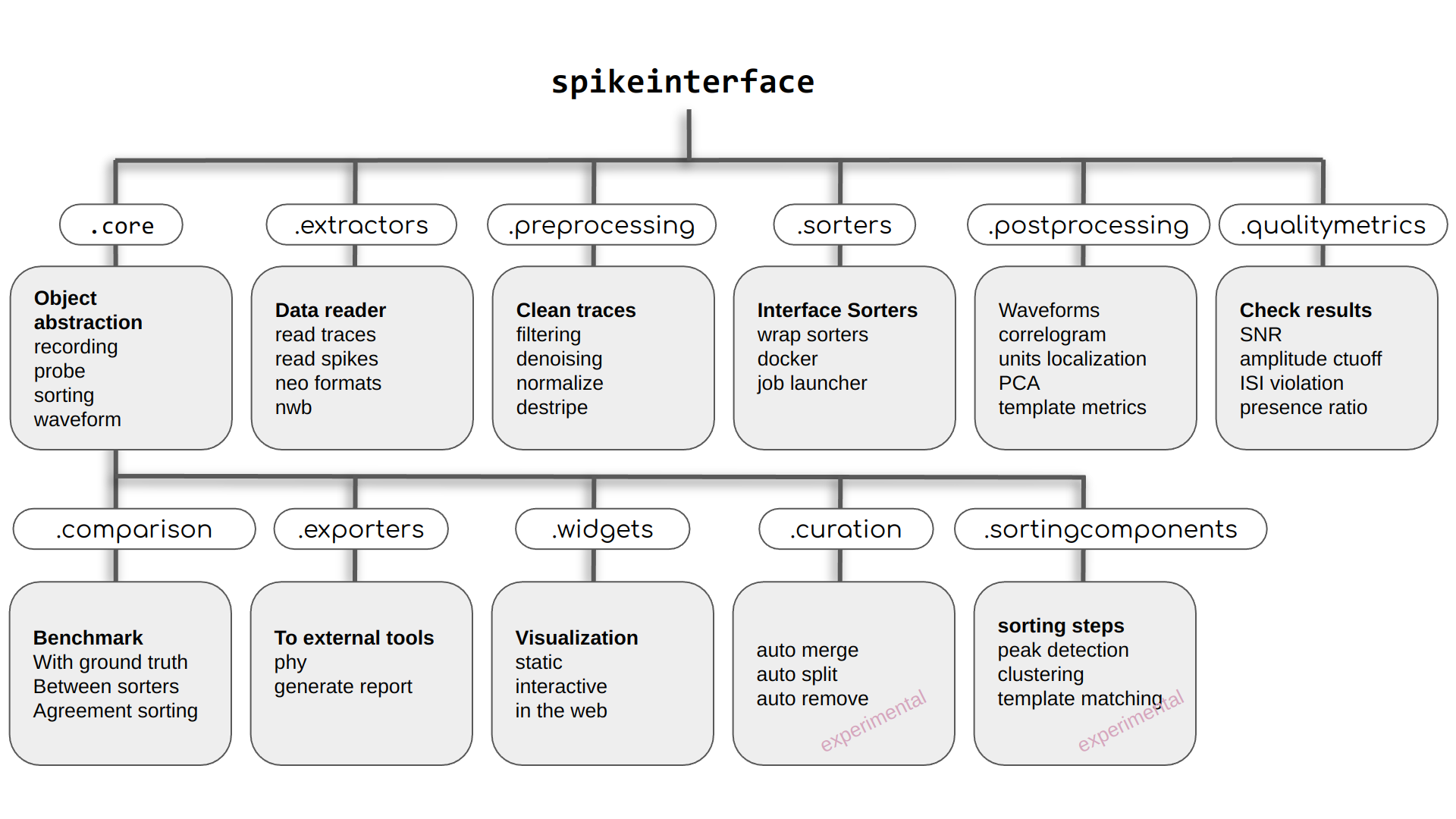
SpikeInterface consists of several sub-packages which encapsulate all steps in a typical spike sorting pipeline:
spikeinterface.curation(under development)spikeinterface.sortingcomponents(under development)