Note
Click here to download the full example code
Preprocessing Tutorial¶
Before spike sorting, you may need to preproccess your signals in order to improve the spike sorting performance.
You can do that in SpikeInterface using the toolkit.preprocessing submodule.
import numpy as np
import matplotlib.pylab as plt
import scipy.signal
import spikeinterface.extractors as se
import spikeinterface.toolkit as st
First, let’s create a toy example:
recording, sorting = se.example_datasets.toy_example(num_channels=4, duration=10, seed=0)
Apply filters¶
Now apply a bandpass filter and a notch filter (separately) to the recording extractor. Filters are also RecordingExtractor objects.
recording_bp = st.preprocessing.bandpass_filter(recording, freq_min=300, freq_max=6000)
recording_notch = st.preprocessing.notch_filter(recording, freq=1000, q=10)
Now let’s plot the power spectrum of non-filtered, bandpass filtered, and notch filtered recordings.
f_raw, p_raw = scipy.signal.welch(recording.get_traces(), fs=recording.get_sampling_frequency())
f_bp, p_bp = scipy.signal.welch(recording_bp.get_traces(), fs=recording.get_sampling_frequency())
f_notch, p_notch = scipy.signal.welch(recording_notch.get_traces(), fs=recording.get_sampling_frequency())
fig, ax = plt.subplots()
ax.semilogy(f_raw, p_raw[0], f_bp, p_bp[0], f_notch, p_notch[0])
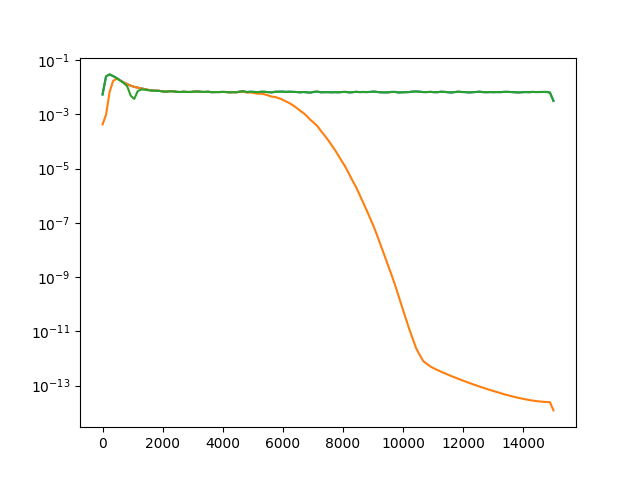
Out:
[<matplotlib.lines.Line2D object at 0x7f3e05894320>, <matplotlib.lines.Line2D object at 0x7f3e0589f080>, <matplotlib.lines.Line2D object at 0x7f3e0589f588>]
Compute LFP and MUA¶
Local field potentials (LFP) are low frequency components of the extracellular recordings. Multi-unit activity (MUA) are rectified and low-pass filtered recordings showing the diffuse spiking activity.
In spiketoolkit, LFP and MUA can be extracted combining the
bandpass_filter, rectify and resample functions. In this
example LFP and MUA are resampled at 1000 Hz.
recording_lfp = st.preprocessing.bandpass_filter(recording, freq_min=1, freq_max=300)
recording_lfp = st.preprocessing.resample(recording_lfp, 1000)
recording_mua = st.preprocessing.resample(st.preprocessing.rectify(recording), 1000)
The toy example data are only contain high frequency components, but these lines of code will work on experimental data
Change reference¶
In many cases, before spike sorting, it is wise to re-reference the signals to reduce the common-mode noise from the recordings.
To re-reference in spiketoolkit you can use the common_reference
function. Both common average reference (CAR) and common median
reference (CMR) can be applied. Moreover, the average/median can be
computed on different groups. Single channels can also be used as
reference.
recording_car = st.preprocessing.common_reference(recording, reference='average')
recording_cmr = st.preprocessing.common_reference(recording, reference='median')
recording_single = st.preprocessing.common_reference(recording, reference='single', ref_channels=[0])
recording_single_groups = st.preprocessing.common_reference(recording, reference='single',
groups=[[0, 1], [2, 3]], ref_channels=[0, 2])
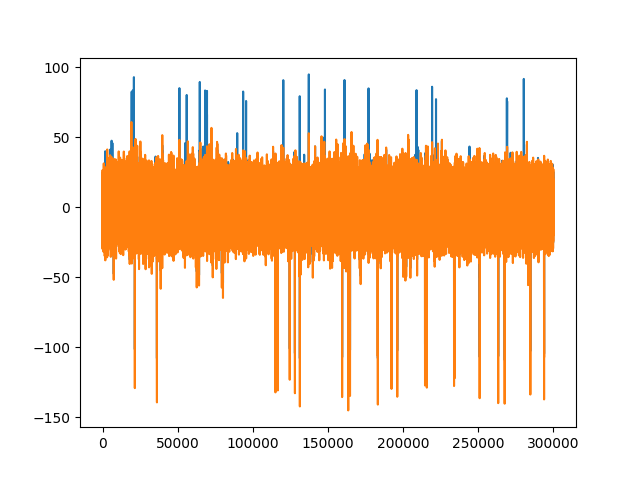
fig1, ax1 = plt.subplots()
ax1.plot(recording_car.get_traces()[0])
ax1.plot(recording_cmr.get_traces()[0])
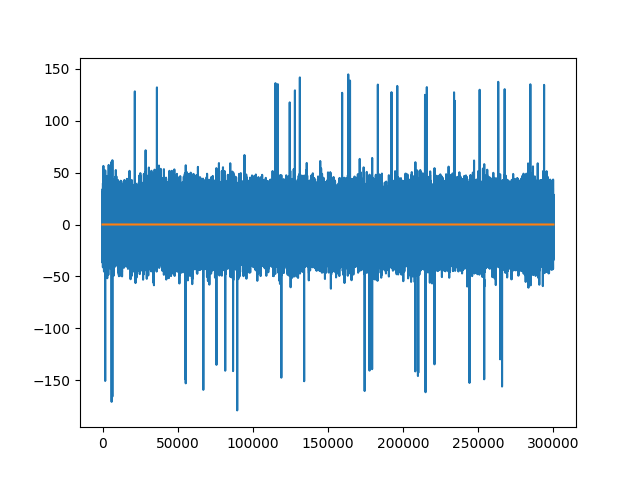
fig2, ax2 = plt.subplots()
ax2.plot(recording_single_groups.get_traces()[1]) # not zero
ax2.plot(recording_single_groups.get_traces()[0])
Out:
[<matplotlib.lines.Line2D object at 0x7f3e056fdb38>]
Remove bad channels¶
In to remove noisy channels from the analysis, the
remove_bad_channels function can be used.
recording_remove_bad = st.preprocessing.remove_bad_channels(recording, bad_channel_ids=[0])
print(recording_remove_bad.get_channel_ids())
Out:
[1, 2, 3]
As expected, channel 0 is removed. Bad channels removal can also be done
automatically. In this case, the channels with a standard deviation
exceeding bad_threshold times the median standard deviation are
removed. The standard deviations are computed on the traces with length
seconds from the middle of the recordings.
recording_remove_bad_auto = st.preprocessing.remove_bad_channels(recording, bad_channel_ids=None, bad_threshold=2,
seconds=2)
print(recording_remove_bad_auto.get_channel_ids())
Out:
[0, 1, 2, 3]
With these simulated recordings, there are no noisy channel.
- Remove stimulation artifacts
In some applications, electrodes are used to electrically stimulate the
tissue, generating a large artifact. In spiketoolkit, the artifact
can be zeroed-out using the remove_artifact function.
# create dummy stimulation triggers
stimulation_trigger_frames = np.array([100000, 500000, 700000])
# large ms_before and s_after are used for plotting only
recording_rmartifact = st.preprocessing.remove_artifacts(recording,
triggers=stimulation_trigger_frames,
ms_before=100, ms_after=200)
fig3, ax3 = plt.subplots()
ax3.plot(recording.get_traces()[0])
ax3.plot(recording_rmartifact.get_traces()[0])
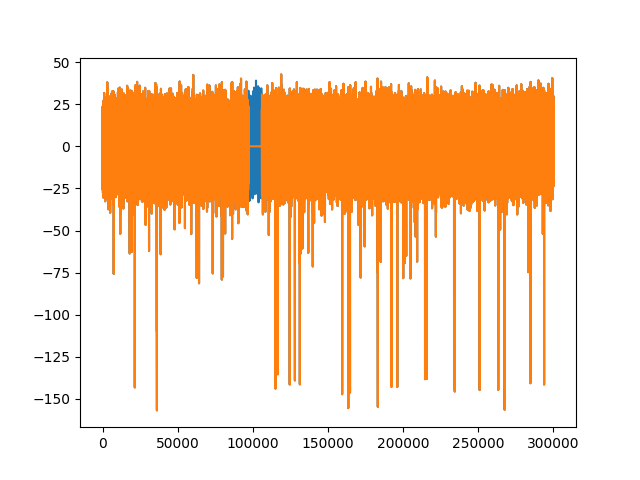
Out:
[<matplotlib.lines.Line2D object at 0x7f3e05684320>]
You can list the available preprocessors with:
print(st.preprocessing.preprocessers_full_list)
Out:
[<class 'spiketoolkit.preprocessing.highpass_filter.HighpassFilterRecording'>, <class 'spiketoolkit.preprocessing.bandpass_filter.BandpassFilterRecording'>, <class 'spiketoolkit.preprocessing.notch_filter.NotchFilterRecording'>, <class 'spiketoolkit.preprocessing.whiten.WhitenRecording'>, <class 'spiketoolkit.preprocessing.common_reference.CommonReferenceRecording'>, <class 'spiketoolkit.preprocessing.resample.ResampleRecording'>, <class 'spiketoolkit.preprocessing.rectify.RectifyRecording'>, <class 'spiketoolkit.preprocessing.remove_artifacts.RemoveArtifactsRecording'>, <class 'spiketoolkit.preprocessing.remove_bad_channels.RemoveBadChannelsRecording'>, <class 'spiketoolkit.preprocessing.transform.TransformRecording'>, <class 'spiketoolkit.preprocessing.normalize_by_quantile.NormalizeByQuantileRecording'>, <class 'spiketoolkit.preprocessing.clip.ClipRecording'>, <class 'spiketoolkit.preprocessing.blank_saturation.BlankSaturationRecording'>, <class 'spiketoolkit.preprocessing.center.CenterRecording'>, <class 'spiketoolkit.preprocessing.mask.MaskRecording'>]
Total running time of the script: ( 0 minutes 2.023 seconds)